Protein structures: Difference between revisions
No edit summary |
No edit summary |
||
| (8 intermediate revisions by the same user not shown) | |||
| Line 2: | Line 2: | ||
{| | {| | ||
|[[image:2e29_asym_r_250.jpg|120px|link=http://www.rcsb.org/pdb/explore.do?structureId=2E29]] | |||
|Ohnishi, S. Pääkkönen, K., Koshiba, S., Tochio, N., Sato, M., Kobayashi, N., Harada, T., Watanabe, S., Muto, Y., Güntert, P., Tanaka, A., Kigawa, T. & Yokoyama, S. Solution structure of the GUCT domain from human RNA helicase II/Guβ reveals the RRM fold, but implausible RNA interactions. [http://dx.doi.org/10.1002/prot.22138 Proteins 74, 133–144 (2009)] | |||
'''PDB [http://www.rcsb.org/pdb/explore.do?structureId=2E29 2E29]''' | |||
|- | |||
|[[image:2jz4_asym_r_250.jpg|120px|link=http://www.rcsb.org/pdb/explore.do?structureId=2JZ4]] | |[[image:2jz4_asym_r_250.jpg|120px|link=http://www.rcsb.org/pdb/explore.do?structureId=2JZ4]] | ||
|Takeda, M., Sugimori, N., Torizawa, T., Terauchi, T., Ono, A. M., Yagi, H., Yamaguchi, Y., Kato, K., Ikeya, T., Güntert, P., Aceti, D. J., Markley, J. L. & Kainosho, M. Structure of the putative 32 kDa myrosinase binding protein from Arabidopsis (At3g16450.1) as determined by the SAIL-NMR method. [http://dx.doi.org/10.1111/j.1742-4658.2008.06717.x FEBS J. 275, 5873–5884 (2008)] | |Takeda, M., Sugimori, N., Torizawa, T., Terauchi, T., Ono, A. M., Yagi, H., Yamaguchi, Y., Kato, K., Ikeya, T., Güntert, P., Aceti, D. J., Markley, J. L. & Kainosho, M. Structure of the putative 32 kDa myrosinase binding protein from Arabidopsis (At3g16450.1) as determined by the SAIL-NMR method. [http://dx.doi.org/10.1111/j.1742-4658.2008.06717.x FEBS J. 275, 5873–5884 (2008)] | ||
'''PDB [http://www.rcsb.org/pdb/explore.do?structureId=2JZ4 2JZ4]''' '''BMRB [http://www.bmrb.wisc.edu./data_library/generate_summary.php?bmrbId= | '''PDB [http://www.rcsb.org/pdb/explore.do?structureId=2JZ4 2JZ4]''' [http://www.rcsb.org/pdb/download/downloadFile.do?fileFormat=NMR&compression=NO&structureId=2JZ4 NMR restraints] '''BMRB [http://www.bmrb.wisc.edu./data_library/generate_summary.php?bmrbId=15607 15607]''' | ||
|- | |- | ||
Latest revision as of 14:53, 9 December 2008
NMR protein structures co-authored by P. Güntert.
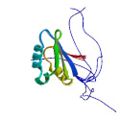
|
Ohnishi, S. Pääkkönen, K., Koshiba, S., Tochio, N., Sato, M., Kobayashi, N., Harada, T., Watanabe, S., Muto, Y., Güntert, P., Tanaka, A., Kigawa, T. & Yokoyama, S. Solution structure of the GUCT domain from human RNA helicase II/Guβ reveals the RRM fold, but implausible RNA interactions. Proteins 74, 133–144 (2009)
PDB 2E29 |
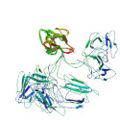
|
Takeda, M., Sugimori, N., Torizawa, T., Terauchi, T., Ono, A. M., Yagi, H., Yamaguchi, Y., Kato, K., Ikeya, T., Güntert, P., Aceti, D. J., Markley, J. L. & Kainosho, M. Structure of the putative 32 kDa myrosinase binding protein from Arabidopsis (At3g16450.1) as determined by the SAIL-NMR method. FEBS J. 275, 5873–5884 (2008)
PDB 2JZ4 NMR restraints BMRB 15607 |
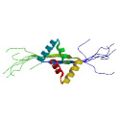
|
Yoshida, H., Furuya, N., Lin, Y. J., Güntert, P., Komano, T. & Kainosho, M. Structural basis of the role of the NikA ribbon-helix-helix domain in initiating bacterial conjugation. J. Mol. Biol. 384, 690–701 (2008) |
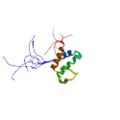
|
Lin, Y. J., Umehara, T., Inoue, M., Saito, K., Kigawa, T., Jang, M. K., Ozato, K., Yokoyama, S., Padmanabhan, B., Güntert, P. Solution structure of the extraterminal domain of the bromodomain-containing protein BRD4. Protein Sci. 17, 2174–2179 (2008) |
|
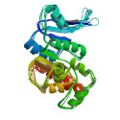
Koglin, A., Löhr, F., Bernhard, F., Rogov, V. R., Frueh, D. P., Strieter, E. R., Mofid, M. R., Güntert, P., Wagner, G., Walsh, C. T., Marahiel, M. A. & Dötsch, V. Structural basis for the selectivity of the external thioesterase of the surfactin-synthetase. Nature 454, 907–911 (2008)
PDB 2RON NMR restraints (SrfTEII) |
|
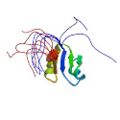
Nagata, T., Suzuki, S., Endo, R., Shirouzu, M., Terada, T., Inoue, M., Kigawa, T, Güntert, P., Hayashizaki, Y., Muto, Y. & Yokoyama, S. The RRM domain of poly(A)-specific ribonuclease (PARN) has a non-canonical binding site for mRNA cap analog recognition. Nucl. Acids Res. 36, 4754–4767 (2008) |
|
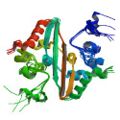
Takeda, M., Chang, C. K., Ikeya, T., Güntert, P., Chang, Y. H., Hsu, Y. L., Huang, T. H. & Kainosho, M. Solution structure of the C-terminal dimerization domain of SARS coronavirus nucleocapsid protein determined by the SAIL-NMR method. J. Mol. Biol. 380, 608–622 (2008) |
|
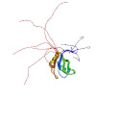
Kuwasako, K., Takahashi, M., Tochio, N., Abe, C., Tsuda, K., Inoue, M., Terada, T., Shirouzu, M., Kobayashi, N., Kigawa, T., Taguchi, S., Tanaka, A., Hayashizaki, Y., Güntert, P., Muto, Y. & Yokoyama, S. Solution structure of the second RNA recognition motif (RRM) domain of murine T cell intracellular antigen-1 (TIA-1) and its RNA recognition mode. Biochemistry 47, 6437–6450 (2008)
PDB 2RNE |
|
Kuwasako, K., Dohmae, N., Inoue, M., Shirouzu, M., Taguchi, S., Güntert, P., Séraphin, B., Muto, Y. & Yokoyama, S. Complex assembly mechanism and an RNA-binding mode of the human p14-SF3b155 spliceosomal protein complex identified by NMR solution structure and functional analyses. Proteins 71, 1617–1636 (2008)
PDB 2FHO NMR restraints |

|
Hwang, E., Ryu, K. S., Pääkkönen, K., Güntert, P., Cheong, H. K., Lim, D. S., Lee, J. O., Jeon, Y. H. & Cheong, C. Structural insight into dimeric interaction of the SARAH domains from Mst1 and RASSF family proteins in the apoptosis pathway. Proc. Natl. Acad. Sci. USA. 104, 9236–9241 (2007)
PDB 2JO8 NMR restraints |

|
Ohnishi, S. Güntert, P., Koshiba, S., Tomizawa, T., Akasaka, R., Tochio, N., Sato, M., Inoue, M., Harada, T., Watanabe, S., Tanaka, T., Shirouzu, M., Kigawa, T. & Yokoyama, S. Solution structure of an atypical WW domain in a novel β-clam-like dimeric form. FEBS Lett. 581, 462–468 (2007)
PDB 2DWV |

|
Kuwasako, K., He, F., Inoue, M., Tanaka, A., Sugano, S., Güntert, P., Muto, Y. & Yokoyama, S. Solution structures of the SURP domains and the subunit-assembly mechanism within the splicing factor SF3a complex in 17S U2 snRNP. Structure 14, 1677-1689 (2006). |

|
ENTH-VHS domain At3g16270(9-135) from Arabidopsis thaliana. López-Méndez, B. & Güntert, P. Automated protein structure determination from NMR spectra. J. Am. Chem. Soc. 128, 13112-13122 (2006). PDB 2DCP NMR restraints Note: Automated structure determination with FLYA. Structure does not supersede the original deposition 1VDY. |

|
Rhodanese homology domain At3g16270(175-295) from Arabidopsis thaliana.
López-Méndez, B. & Güntert, P. Automated protein structure determination from NMR spectra. J. Am. Chem. Soc. 128, 13112-13122 (2006). PDB 2DCQ NMR restraints Note: Automated structure determination with FLYA. Structure does not supersede the original deposition 1VEE. |

|
Src homology domain 2 from the human feline sarcoma oncogene Fes.
López-Méndez, B. & Güntert, P. Automated protein structure determination from NMR spectra. J. Am. Chem. Soc. 128, 13112-13122 (2006). |

|
NikA N-terminal fragment.
Yoshida, H., Furuya, N., Lin, Y. J., Güntert, P., Komano, T. & Kainosho, M. PDB 2BA3 |

|
Jurt, S., Aemissegger, A., Güntert, P., Zerbe, O. & Hilvert, D. A photoswitchable miniprotein based on the sequence of avian pancreatic polypeptide. Angew. Chem. Int. Ed. 45, 6297-6300 (2006).
PDB 2H4B NMR restraints (cis-1-PP) |

|
Hamada, T., Ito, Y., Abe, T., Hayashi, F., Güntert, P., Inoue, M., Kigawa, T., Terada, T., Shirouzu, M., Yoshida, M., Tanaka, A., Sugano, S., Yokoyama, S. & Hirota, H. Solution structure of the antifreeze-like domain of human sialic acid synthase. Protein Sci. 15, 1010-1016 (2006).
PDB 1WVO NMR restraints |

|
Aachmann, F. L., Svanem, B. I. G., Güntert, P., Petersen, S. B., Valla, S. & Wimmer, R. NMR structure of the R-module - A parallel β-roll subunit from an Azotobacter vinelandii mannuronan C-5 epimerase. J. Biol. Chem. 281, 7350-7356 (2006).
PDB 2AGM NMR restraints |

|
Stereo-array isotope labeled (SAIL) maltodextrin-binding protein MBP.
Kainosho, M., Torizawa, T., Iwashita, Y., Terauchi, T., Ono, A. M. & Güntert, P. Optimal isotope labelling for NMR protein structure determinations. Nature 440, 52-57 (2006). PDB 2D21 BMRB 6807 |

|
Stereo-array isotope labeled (SAIL) calmodulin.
Kainosho, M., Torizawa, T., Iwashita, Y., Terauchi, T., Ono, A. M. & Güntert, P. Optimal isotope labelling for NMR protein structure determinations. Nature 440, 52-27 (2006). PDB 1X02 BMRB 6541 |

|
Li, H., Inoue, M., Yabuki, T., Aoki, M., Seki, E., Matsuda, T., Nunokawa, E., Motoda, Y., Kobayashi, A., Terada, T., Shirouzu, M., Koshiba, S., Lin, Y. J., Güntert, P., Suzuki, H., Hayashizaki, Y., Kigawa, T. & Yokoyama, S. Solution structure of the mouse enhancer of rudimentary protein reveals a novel fold. J. Biomol. NMR 32, 329-334 (2005).
PDB 1WWQ |

|
Scott, A., Pantoja-Uceda, D., Koshiba, S., Inoue, M., Kigawa, T., Terada, T., Shirouzu, M., Tanaka, A., Sugano, S., Yokoyama, S. & Güntert, P. Solution structure of the Src homology 2 domain from the human feline sarcoma oncogene Fes. J. Biomol. NMR 31, 357-361 (2005).
PDB 1WQU BMRB 6331 |

|
Nameki, N., Tochio, N., Koshiba, S., Tochio, N., Inoue, M., Yabuki, T., Aoki, M., Seki, E., Matsuda, T., Saito, M., Ikari, M., Watanabe, M., Terada, T., Shirouzu, M., Yoshida, M., Hirota, H., Tanaka, A., Hayashizaki, Y., Güntert, P., Kigawa, T. & Yokoyama, S. Solution structure of the PWWP domain of the hepatoma-derived growth factor family. Prot. Sci. 14, 756-764 (2005).
PDB 1N27 |

|
Calzolai, L., Lysek, D. A., Pérez, D. R., Güntert, P. & Wüthrich, K. Prion protein NMR structures of chicken, turtle and frog. Proc. Natl. Acad. Sci. USA 102, 651-655 (2005).
PDB 1U3M NMR restraints (chicken PrP(121-225)) 1U5L NMR restraints (turtle PrP(121-225)) 1XU0 NMR restraints (frog PrP(90-222)) BMRB 6269 (chicken PrP(128-242)) 6270 (chicken PrP(25-242)) 6282 (turtle PrP(121-225)) |

|
Lysek, D. A., Schorn, C., Nivon, L. G., Esteve-Moya, V., Christen, B., Calzolai, L., von Schroetter, C., Fiorito, F., Herrmann, T., Güntert, P. & Wüthrich, K. Prion protein NMR structures of cat, dog, pig and sheep. Proc. Natl. Acad. Sci. USA 102, 640-645 (2005).
PDB 1XYJ NMR restraints (cat PrP(121-231)) 1XYK NMR restraints (dog PrP(121-231)) 1XYQ NMR restraints (pig PrP(121-231)) 1XYU NMR restraints (sheep PrP[H168](121-231)) 1Y2S NMR restraints (sheep PrP[R168](121-231)) BMRB 6377 (cat PrP(121-231)) 6378 (dog PrP(121-231)) 6380 (pig PrP(121-231)) 6381 (sheep PrP[H168](121-231)) 6403 (sheep PrP[H168](121-231)) |

|
Pääkkönen, K., Tossavainen, H., Permi, P., Rakkolainen, H., Rauvala, H., Raulo, E., Kilpeläinen, I. & Güntert, P. Solution structures of the first and fourth TSR domains of F-spondin. Proteins 64, 665-672 (2006).
PDB 1SZL (TSR1) 1VEX (TSR4)
PDB 1VDY BMRB 5928 |

|
Iwai, H., Forrer, P., Plückthun, A. & Güntert, P. NMR solution structure of the monomeric form of the bacteriophage λ capsid stabilizing protein gpD. J. Biomol. NMR 31, 351-356 (2005).
PDB 1VD0 BMRB 4936 |

|
Pantoja-Uceda, D., López-Méndez, B., Koshiba, S., Inoue, M., Kigawa, T., Terada, T., Shirouzu, M., Tanaka, A., Seki, M., Shinozaki, K., Yokoyama, S. & Güntert, P. Solution structure of the rhodanese homology domain At4g01050(175-295) from Arabidopsis thaliana. Prot. Sci. 14, 224-230 (2005).
PDB 1VEE BMRB 5929 |

|
Nameki, N., Yoneyama, M., Koshiba, S., Tochio, N., Inoue, M., Seki, E., Matsuda, T., Tomo, Y., Harada, T., Saito, K., Kobayashi, N., Yabuki, T., Aoki, M., Nunokawa, E., Matsuda, N., Sakagami, N., Terada, T., Shirouzu, M., Yoshida, M., Hirota, H., Osanai, T., Tanaka, A., Arakawa, T., Carninci, P., Kawai, J., Hayashizaki, Y., Kinoshita, K., Güntert, P., Kigawa, T. & Yokoyama, S. Solution structure of the RWD domain of the mouse GCN2 protein. Prot. Sci. 13, 2089-2100 (2004).
PDB 1UKX |

|
Fernández, C., Hilty, C., Wider, G., Güntert, P. & Wüthrich, K. NMR Structure of the integral membrane protein OmpX. J. Mol. Biol. 336, 1211-1221 (2004).
PDB 1Q9G BMRB 4936 |

|
FHA domain of mouse afadin 6.
PDB 1WLN |

|
Vanwetswinkel, S., Kriek, J., Andersen, G. R., Güntert, P., Dijk, J., Canters, G. W. & Siegal, G. Solution structure of the 162 residue C-terminal domain of the human elongation factor 1Bγ. J. Biol. Chem. 278, 43443-43451 (2003).
PDB 1PBU NMR restraints BMRB 5628 |

|
Hilge, M., Siegal, G., Vuister, G. W., Güntert, P., Gloor, S. M. & Abrahams, J. P. ATP-induced conformational changes of the nucleotide-binding domain of Na,K-ATPase. Nat. Struct. Biol. 10, 468-474 (2003).
PDB 1MO7 (free form) 1MO8 (ATP-bound form) BMRB 5577 (free form) 5576 (ATP-bound form) |

|
Lührs, T., Riek, R., Güntert, P. & Wüthrich, K. NMR structure of the human doppel protein. J. Mol. Biol. 326, 1549-1557 (2003).
PDB 1LG4 NMR restraints BMRB 5145 |

|
Zahn, R., Güntert, P., von Schroetter, C. & Wüthrich, K. NMR structure of a human prion protein with two disulfide bridges. J. Mol. Biol. 326, 225-234 (2003).
PDB 1H0L NMR restraints BMRB 5378 |

|
Enggist, E., Thöny-Meyer, L., Güntert, P. & Pervushin, K. NMR structure of the heme chaperone CcmE reveals a novel functional motif. Structure 10, 1551-1557 (2002).
PDB 1SR3 NMR restraints |

|
Lee, D., Damberger, F. D., Peng, G., Horst, R., Güntert, P., Nikonova, L., Leal, W. S. & Wüthrich, K. NMR structure of the unliganded Bombyx mori pheromone-binding protein at physiological pH. FEBS Lett. 531, 314-318 (2002).
PDB 1LS8 |

|
Ellgaard, L., Bettendorff, P., Braun, D., Herrmann, T., Fiorito, F., Jelesarov, I., Herrmann, T., Güntert, P., Helenius, A. & Wüthrich, K. NMR structures of 36 and 73-residue fragments of the calreticulin P-domain. J. Mol. Biol. 322, 773-784 (2002).
PDB 1K9C NMR restraints (residues 189-261) 1K91 (residues 221-256) BMRB 5204 (residues 189-261) 5205 (residues 221-256) |

|
Miura, T., Klaus, W., Ross, A., Güntert, P. & Senn, H. The NMR structure of the class I human ubiquitin-conjugating enzyme 2b. J. Biomol. NMR 22, 89-92 (2002).
PDB 1JAS NMR restraints BMRB 5038 |

|
Horst, R., Damberger, F., Luginbühl, P., Güntert, P., Peng, G., Nikonova, L., Leal, W. S. & Wüthrich, K. NMR structure reveals intramolecular regulation mechanism for pheromone binding and release. Proc. Natl. Acad. Sci. USA 98, 14374-14379 (2001).
PDB 1GM0 NMR restraints BMRB 4849 |

|
Ellgaard, L., Riek, R., Herrmann, T., Güntert, P., Braun, D., Helenius, A. & Wüthrich, K. NMR structure of the calreticulin P-domain. Proc. Natl. Acad. Sci. USA 98, 3133-3138 (2001).
PDB 1HHN BMRB 4878 |

|
Calzolai, L., Lysek, D. A., Güntert, P., von Schroetter, C., Riek, R., Zahn, R. & Wüthrich, K. NMR structures of three single-residue variants of the human prion protein. Proc. Natl. Acad. Sci. USA 97, 8340-8345 (2000).
PDB 1E1G NMR restraints (variant M166V) 1E1P NMR restraints (variant S170N) 1E1U NMR restraints (variant R220K) BMRB 4736 (variant M166V) 4620 (variant R220K) |

|
Riek, R., Prêcheur, P., Wang, Y., Mackay, E. A., Wider, G., Güntert, P., Liu, A., Kägi, J. H. R. & Wüthrich, K. NMR structure of the sea urchin (Strongylocentrotus purpuratus) metallothionein MTA. J. Mol. Biol. 291, 417-428 (1999).
PDB 1QJK NMR restraints (alpha domain) 1QJL NMR restraints (beta domain) BMRB 4363 |

|
Pellecchia, M., Güntert, P., Glockshuber, R. & Wüthrich, K. The NMR solution structure of the periplasmic chaperone FimC. Nature Struct. Biol. 5, 885-890 (1998).
PDB 1BF8 BMRB 4070 |

|
Ottiger, M., Zerbe, O., Güntert, P. & Wüthrich, K. The NMR solution conformation of unligated human cyclophilin A. J. Mol. Biol. 272, 64-81 (1997).
PDB 1OCA NMR restraints |

|
Antuch, W., Güntert, P. & Wüthrich, K. Ancestral beta gamma-crystallin precursor structure in a yeast killer toxin, Nature Struct. Biol. 3, 662-665 (1996).
PDB 1WKT NMR restraints BMRB 5255 |

|
Antuch, W., Güntert, P., Billeter, M., Hawthorne, T., Grossenbacher, H. & Wüthrich, K. NMR solution structure of the recombinant tick anticoagulant protein (rTAP), a factor Xa inhibitor from the tick Ornithodoros moubata. FEBS Lett. 352, 251-257 (1994).
PDB 1TAP |

|
Berndt, K. D., Güntert, P. & Wüthrich, K. The NMR solution structure of dendrotoxin K from the venom of Dendroaspis polylepis polylepis. J. Mol. Biol. 234, 735-750 (1993).
PDB 1DTK NMR restraints |

|
Szyperski, T., Güntert, P., Stone, S. R. & Wüthrich, K. The NMR solution structure of hirudin(1-51) and comparison with corresponding three-dimensional structures determined using the complete 65-residue hirudin polypeptide chain. J. Mol. Biol. 228, 1193-1205 (1992).
PDB 1HIC NMR restraints |

|
Berndt, K. D., Güntert, P., Orbons, L. P. M. & Wüthrich, K. Determination of a high-quality NMR solution structure of the bovine pancreatic trypsin inhibitor (BPTI) and comparison with three crystal structures. J. Mol. Biol. 227, 757-775 (1992).
PDB 1PIT NMR restraints |

|
Güntert, P., Qian, Y. Q., Otting, G., Müller, M., Gehring, W. J. & Wüthrich K. Structure determination of the Antp(C39->S) homeodomain from nuclear magnetic resonance data in solution using a novel strategy for the structure calculation with the programs DIANA, CALIBA, HABAS and GLOMSA. J. Mol. Biol. 217, 531-540 (1991).
PDB 2HOA |